Science’s efforts towards understanding life have reached a remarkably satisfying point, which earlier researchers could only have dreamed of.
Scientists’ understanding of life remains peculiarly unsatisfying, despite our best efforts.
Both these declarations are true, to judge by a recent conference surveying the state of biology. The meeting, held in Dublin last September to mark the 75th anniversary of physicist Erwin Schrödinger’s speculative book What is Life?, looked to the future as well as celebrating the past. And it suggested that the tension between these verdicts may energise the most interesting work to come.
The achievements of the approach that has dominated the last three-quarters of a century – taking organisms apart to inspect all the bits – surely are remarkable. Schrödinger, with his prescient emphasis in 1943 that the molecular information store underlying heredity was probably an “aperiodic crystal”, helped to inspire one of the two main research streams of molecular biology. That led from the solution of DNA’s structure a decade later to the completion of the entire sequence of our own DNA in the human genome project around the end of the millennium.
A second stream has had equally spectacular success. A living cell is packed with intricate molecular machines, and we can now dissect their workings in as much detail as we desire. Take the ribosome, which makes new chains of amino acids according to the order specified by the sequence of bases in a piece of RNA, the other nucleic acid that cells use to carry information. These could once be seen only as vague blobs in an electron micrograph. But, as the Israeli crystallographer Ada Yonath reminded the Dublin audience, we now know in minute detail (thanks in no small part to her own work, for which she won the 2009 Nobel Prize in Chemistry) the structure of this complex assemblage of structural RNA, accessorised with more than 50 smaller proteins.
More, we know exactly how the parts move and interact. There are high-res animations on YouTube showing how they join the right amino acids together, rapidly and efficiently, in the right order, and release them as a newly synthesised protein. The animation is designed to look like a machine on a production line, not part of the swirl of molecules that wash around inside cells. Still, it’s impossible not to watch this elegantly choreographed microscopic dance, repeated millions of times in every cell, without feeling that you are seeing something fundamental to all life.
The ribosome is emblematic of the fact that there is no component of a cell, no matter how intricate, that we cannot now isolate from the wider organism and understand down to the very atoms if we apply the tools to hand. We are able to modify these tiny biological fragments, too – chemically or by genetic manipulation – and predict more or less what the effect will be.
However, that doesn’t mean we know what will happen when we put them back into a cell, an organ or a whole creature. And it is these higher-level biological phenomena that the discipline is ultimately interested in understanding.

So what do biologists do when the number of variables, interactions between components or possible arrangements of parts increases by a few orders of magnitude – when, as University of California, San Francisco neuroscientist Saul Kato put it, you can’t work out how something is working just by looking at it?
There are no clear answers. At the moment, discussion mainly yields alternative ways of framing the same question. For example, what do we do with the data accumulating from new techniques in genetics, cell biology or neuroscience? The data mountain is itself a result of past successes: databanks are filling at an ever-increasing rate with DNA sequences, protein structures and detailed measurements of changing levels of smaller molecules. In fact, Kato suggested, biology is in an age of excessive data. They can often be displayed prettily, in lots of different ways, but they mostly elude interpretation.
As Kato also emphasised, outlining networks doesn’t give many clues about how they work. There is ambitious talk of mapping all the neural circuits in a brain, perhaps even a human brain – building a “connectome” to match the sequencing of the human genome. But we have had access to the entire, fixed, connectome of one simple organism, the nematode worm Caenorhabditis elegans, for 30 years now. And it turns out that having a complete wiring diagram of its few hundred neurons still leaves many unanswered questions about its behaviour.
Kato suggests that the answers lie in “computational biology”, perhaps of the kind exhibited by Danielle Bassett of the University of Pennsylvania, who applies mathematical techniques of network analysis to the dynamics of neural circuitry, among many other things. The results are still somewhat abstract, though, and remain some way from the other goal many speakers reach for when discussing these problems: developing something called “systems biology”.
That’s another catch-all term that needs a lot of unpacking, but one of its key features emerged from several talks. For University of California, Santa Barbara cognitive neuroscientist Michael Gazzaniga, famed for his experiments with “split brain” patients, it means taking account of an overall architecture of an organism. Thinking about the gap between neurons and minds encourages the belief that life is a layered system, and each layer has an inbuilt vocabulary appropriate for its own operation. But, Gazzaniga added, how the layers communicate remains unknown.
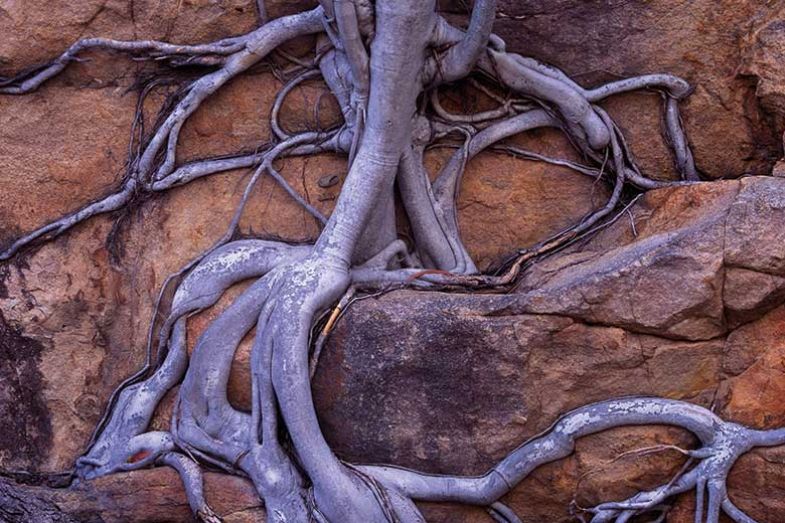
Ottoline Leyser of the University of Cambridge, who studies the ostensibly less complex world of plants, made a similar point about the increasingly evident gaps in our knowledge of whole organisms. The defining feature of biology during the past few decades has been figuring out details of the parts, she said. But “biological systems don’t think they have parts”.
What they do have are systems that are dynamic, and that show self-organisation across scales that span many orders of magnitude. We know that inputs at one level of a system can produce responses at all the others. But biologists are still trying to find the vocabulary, or the intellectual toolkit, for understanding the kind of self-organisation that leads a plant, for example, to adjust its branching pattern to deal with a change in its surroundings or its nutrient supply. Leyser’s big question is: “How does information flow across scales?”
This is all very reminiscent of Schrödinger in one respect. He proposed that deep understanding of organisms would require some new principles, perhaps even new laws of physics. That part of his book never really found any serious adherents. But it does now look as if, after three-quarters of a century spent becoming virtuosi of molecular dissection, biologists are going to need genuinely new ideas to take their understanding of living systems to the next level. On the evidence of the Dublin meeting, they aren’t quite there yet.
Jon Turney is a science writer and an associate lecturer at Bath Spa University.
Our physical nature: Schrödinger’s catch
The question “What is life?” triggers idiosyncratic answers. In the context of Schrödinger’s book of that name, the answers focus on the place of the laws of physics in biology.
For many biologists and physicists alike – all of whom are human, after all – the suggestion that physics might explain humanity seems to trigger angst about human nature, carrying implications for our understanding of consciousness, creativity and free will.
I am not disturbed. I like thinking. Can my thoughts be explained through physics? Of course they can: they are physically encoded. Does that make me any less human? No – why should it? Thinking is no less joyful and fulfilling to me just because it has a physical basis. Schrödinger’s description of life as a dynamic system far from equilibrium, maintained by a constant supply of negative entropy (food), is for me inspiring, not deflating.
Schrödinger’s treatise focuses on the physical nature of genes, inferred from their behaviour in various biological experiments. He provides a reasonable estimate of gene size, and famously describes them as aperiodic crystals. In the intervening 75 years, we have discovered much about genes. But through this work, it is clear that genes do not really have a simple physical definition. They have a basis in physics, but the value of the concept of a gene encapsulates far more than the bit of DNA that can be delineated as carrying the information needed to deliver a particular biochemical function.
A gene is a unit of heredity. The central experiments that underpinned both Schrödinger’s deductions and the coining of the word “gene” relate to the inheritance of organismal characteristics – the wrinkliness of peas, the colour of fruit flies’ eyes or a person’s ability to clot blood when they are cut. For these examples, there happens to be a relatively simple linear series of events linking DNA, via RNA and protein, to a particular biochemical reaction that directly underpins those properties. But many traits, such as height, are complex, underpinned by many genes – and by interactions between those genes and the environment.
For me, as a developmental biologist, the most interesting characteristic of an organism is how it got to be an organism in the first place. How does a single cell, the fertilised egg, become a multi-cellular organism with many different cell types in the right relative arrangement, working together to make a functioning organism? How are peas, flies’ eyes and blood made?
For these questions, the relationship between genes and organisms is very poorly understood. Genes are switched off and on to drive protein production in patterns across space and time. These patterns feed back to change which genes are off and on, and feed forward to change the properties of cells, tissues and organs – which, in turn, feed back to influence the behaviour of tissues, cells and genes. Across biological scales, highly dynamic and interconnected processes operate in diverse ways to orchestrate the construction, maintenance and operation of organisms, far from equilibrium. It’s astonishing.
We have some understanding of the molecular-scale events that characterise life, and the organismal-scale properties that emerge, but the relationship between them is still poorly understood. We need new ways of thinking about the dynamic connections between the scales of organisation in biology, so that we can bridge this gap in understanding. It is perhaps because I am particularly interested in this gap, and the mind-boggling things it can do, that I am not in any way dismayed by the idea that the answers have a physical basis. For me, that’s breathtakingly beautiful, not crushingly demystifying.
Ottoline Leyser is director of the Sainsbury Laboratory, University of Cambridge.
Register to continue
Why register?
- Registration is free and only takes a moment
- Once registered, you can read 3 articles a month
- Sign up for our newsletter
Subscribe
Or subscribe for unlimited access to:
- Unlimited access to news, views, insights & reviews
- Digital editions
- Digital access to THE’s university and college rankings analysis
Already registered or a current subscriber?